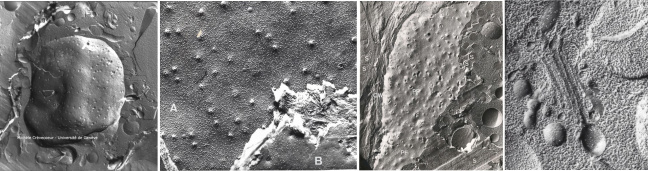
Cryofracture
The cryofracture technique consists in rapid freezing of a biological sample and its fracturing at low temperature under vacuum. A replica of the fractured surface is then generated by evaporation of carbon and platinum. The replica is separated from the tissue leaving a thin Pt – Carbon film observed in a transmission electron microscope (TEM).
Preparation of samples
The biological sample is submitted to rapid/ultrarapid freezing. When the temperature decreases below the freezing point the general hydration water of the living matter crystallizes in situ in many small crystals. These may damage samples, in particular the cellular membranes.
A high speed of freezing partly avoids the formation of these crystals. It is also possible to protect the sample before freezing by a pretreatment with a cryoprotectant such as an aqueous glycerol (25%). And to prevent plasmolysis of cells resulting from glycerol treatment the sample may be fixed with glutaraldehyde.
Sample preparation is function of water content of the tissues. In general, for samples with water content varying from 50 to 80% a prefixation with 4% glutaraldehyde and 25% glycerol treatment before freezing result in nice images. For samples with very low water content as dry ungerminated seeds (5 to 10 % water content) or seeds germinated for a few hours (20 to 45% water content) higher concentrations of both glycerol and glutaraldehyde (50%) result in good qualities images.
Fracture of frozen samples : on the right
The frozen samples on their support are disposed in a particular instrument (example Balzers BA 360 M) at -100°C under high vacuum. They are fractured with a microtome equipped with Eversharp – shick blade and maintained at low temperature with liquid nitrogen.
Depending on the level of fracture it is possible to image the outer surface of sample or internal structures of the sample. However, the technique is extremely useful for examining the macromolecular structure of membranes. Indeed, the fracture preferentially splits the membranes along weak hydrophobic planes and separates membrane phospholipids into two opposite and complementary faces (see figure on the right).
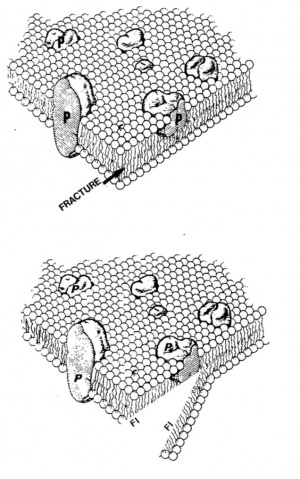
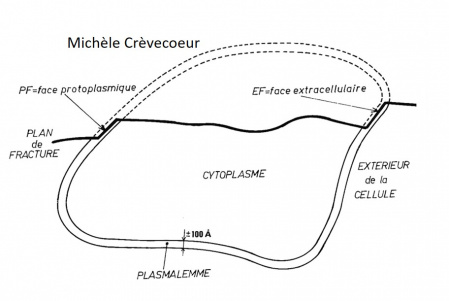
The fracture occurs inside the phospholipid bilayer (conituous line). The process yields two fracture faces designated according to the nomenclature of Branton et al. (1975): the PF face is half of the membrane that remains attached to the cytoplasm (protoplasmic face) and the EF face, half of the bimolecular leaflet that remains attached to the extracellular medium (cell wall in plant cells) after fracturing.
Shadowing
The details of structures revealed by the fracture are covered at -100 °C by a thin layer of carbon – Pt evaporated over the fractured surface. Carbon and platinum are evaporated from an angle (30 to 45°) that confers a three-dimensional image of the fractured surface. A thin layer of carbon is then evaporated to stabilize and reinforce the replica. Both layers, Pt – C and C form the replica.
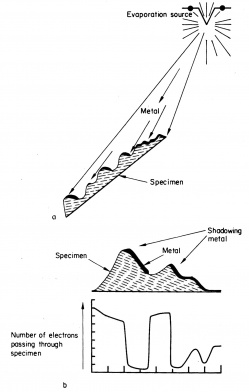
The structures of the specimen exposed to the source will be covered by carbon – Pt will stop electrons and will appear electron dense. On the contrary the structural details of the surface that are on the opposite of the source will not be covered by Pt – C. They will allow the passage of electrons and they will appear bright.
On the right : principle of shadowing according to (1970) in « Electron Microscopy and plant ultrastructure ». Ed McGraww Hill.
a: evaporation of metal from a filament.
b: proportion of electrons that will reach the screen of the transmission electron microscope.
Cleaning of the replica
The vacuum is broken, and the sample covered with the replica is removed. It is firstly immersed in a solution of 10% sulfuric acid then in 75% sulfuric acid to digest the biological material adhering to the replica. Once the digestion is complete the replica is transferred in bi distilled – water to be washed several times. It is finally mounted on formvar copper coated electron microscope grid and observed with a transmission electron microscope.
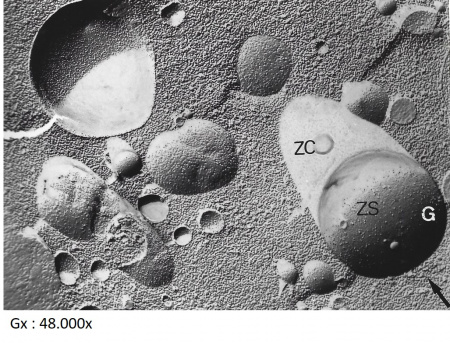
Beow portions of the plasma membrane faces with small globular structures corresponding to intramembrane proteins. Their projected shadow appears bright.

